BIOLOGICS CHARACTERIZATION SERVICES
Mass spectrometry (MS) is widely used in the characterization of biomolecules including peptide and protein therapeutics. These biotechnology products have seen rapid growth over the past few decades and continue to dominate the global pharmaceutical market. Advances in MS instrumentation and techniques have enhanced protein characterization capabilities and supported an increased development of biopharmaceutical products.
Intact Mass Analysis
Intact mass analysis is the assessment of a protein’s total molecular weight by mass spectrometry (MS) without prior digestion or fragmentation of the molecule of interest. Molecular weight determination forms part of the ICH Q6B guidelines for physicochemical analysis of biological products: what use is this information in the analytical characterization of biopharmaceuticals?
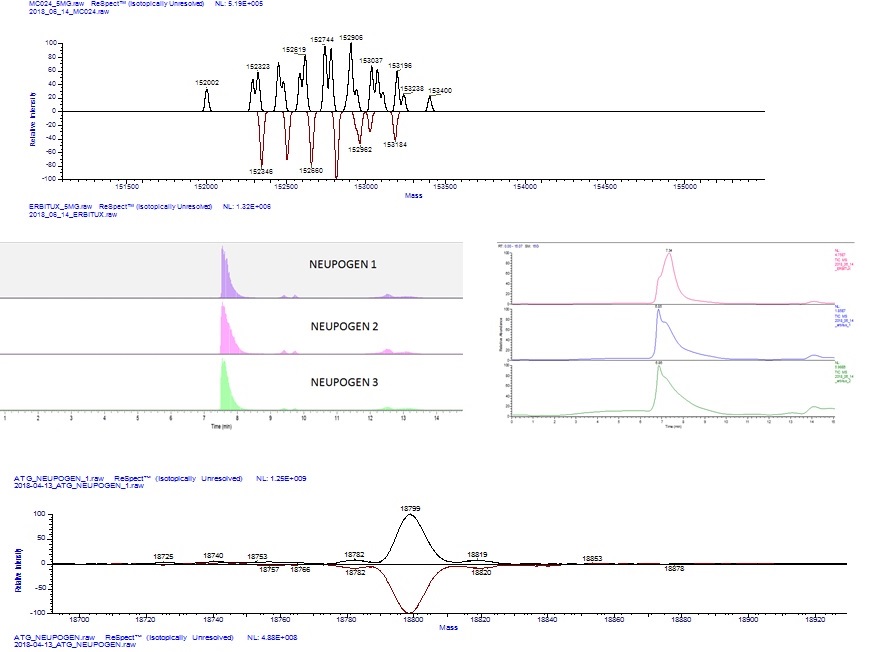
High resolution MS allows the average molecular weight of large proteins, for example monoclonal antibodies (mAbs), to be measured with accuracy better than ± 0.01%. The observed mass can then be compared to the expected mass for a given amino acid sequence.
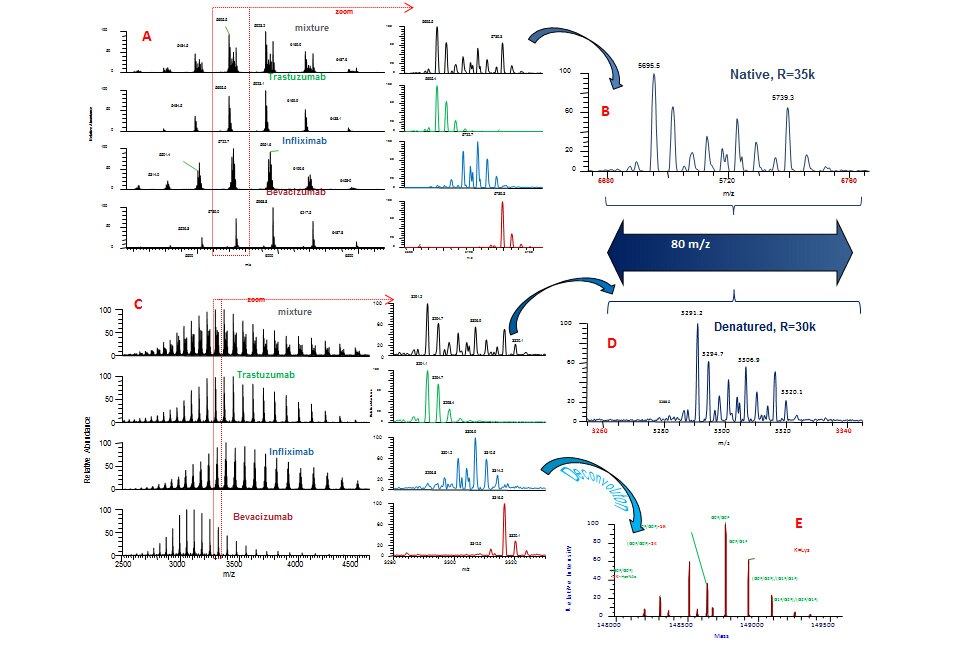
Peptide Mapping
Peptide mapping is a critical workflow in biotherapeutic protein characterization and is essential for elucidating the primary amino acid structure of proteins. For recombinant protein pharmaceuticals, such as monoclonal antibodies (mAbs) and antibody-drug conjugates (ADCs), peptide mapping is used for proof of identity, primary structural characterization and quality assurance/quality control (QA/QC). Global regulatory agencies, including US Food and Drug Administration (US FDA) and European Medicines Agency (EMA), look to harmonized guidelines from the International Council for Harmonisation (ICH). ICH Q6B covers the test procedures and acceptance criteria for biologic drug products, and specifies the use of peptide mapping as a critical quality test procedure for drug characterization used to confirm desired product structure for lot release purposes.
In order to generate a peptide map, the therapeutic protein must first be digested into its constituent peptides via a chemical or enzymatic reaction. Robust separation and identification of the resultant peptides then provides insight into a protein’s full sequence information; displaying each amino acid component and the surrounding amino acid microenvironment, including disulfide linkage information. Structural characterization at this level highlights post translational modifications (PTMs) such as site-specific glycosylation, amino acid substitutions (sequence variants) and/or truncations which may result from erroneous transcription of complementary DNA. Within a bioproduction environment, peptide mapping is necessary for manufacturing process monitoring and QC. It facilitates product comparability testing, which is necessary to identify any product-related impurities, such as deamidation and/or oxidation following any formulation, manufacturing process or storage change.
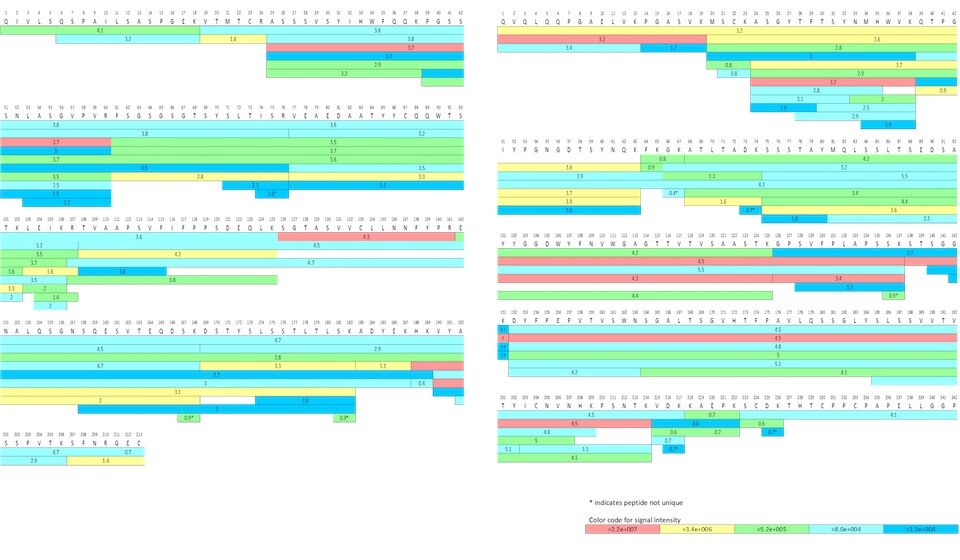
Due to its complexity and inherent variability, peptide mapping is generally performed in a comparative manner; for example, biosimilars would be compared to a reference or control substance, such as the innovator biologic, in a side-by-side experiment. An in-depth analysis is then required to identify minor and even isobaric differences in protein primary structure.
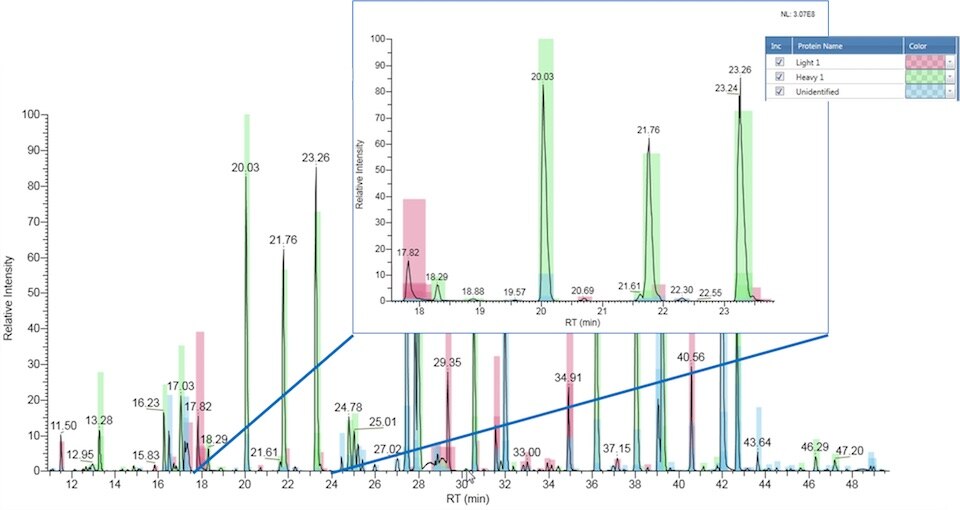
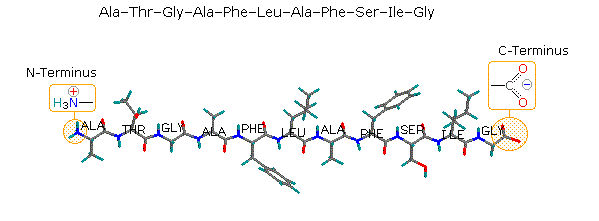
N-terminal and C- terminal sequencing
N-terminal sequencing is widely accepted as a reliable tool for protein characterization throughout all stages of drug discovery and biopharmaceutical manufacturing. The two major direct methods of protein N-terminal sequencing are Edman degradation and mass spectrometry. A strategic approach to the enzyme digestion via an in silico digest selectively optimises the N-terminal peptide fragmentation to achieve suitable peptides of 20-50 amino acid residues.The intact peptide is then deliberately fragmented within a mass spectrometer in order to gain structural information from the fragment ions
Applications:
- Confirmation of structure
- Demonstration of comparability
- Modified terminals (including acetylation and pyroglutamic acids)
- Degradation of proteins (perhaps due to deamidation)
- Determination of enzymatic cleavage specificity